1.24.4 Viewing Alignment Sequences
Alignments are visible in the DNA view as well as the circular and linear view. Both the plus and minus strands of the DNA template are searched for sequence alignments.
DNA View
- Each alignment is displayed in a continuous row, unless overlapping, whereby the alignment is displayed as an additional row (Figure 1.24.4.1).
- Trimmed regions are not deleted, but are shown as greyed out areas of sequence.
Grey bars above the DNA template sequence represent the alignment score. Each bar represents a base - the height of the bar depends on how many correct alignment hits there are. Each correct alignment hit receives a score of 1. For example, if there are 3/3 correct alignments, the alignment score is 3 and the height of the bar is 3x. (Coming soon)
The alignment sequences each have three layers:
- The chromatogram curves (optional layer): blue=C, red=T, black=G, green= A.
- The chromatogram sequence: DNA sequence as in the chromatogram file
- The alignment axis: Axis according to the alignment in the alignment file
Behind the chromatogram data lays the confidence scores – hovering over them reveals the exact percentage (coming soon).
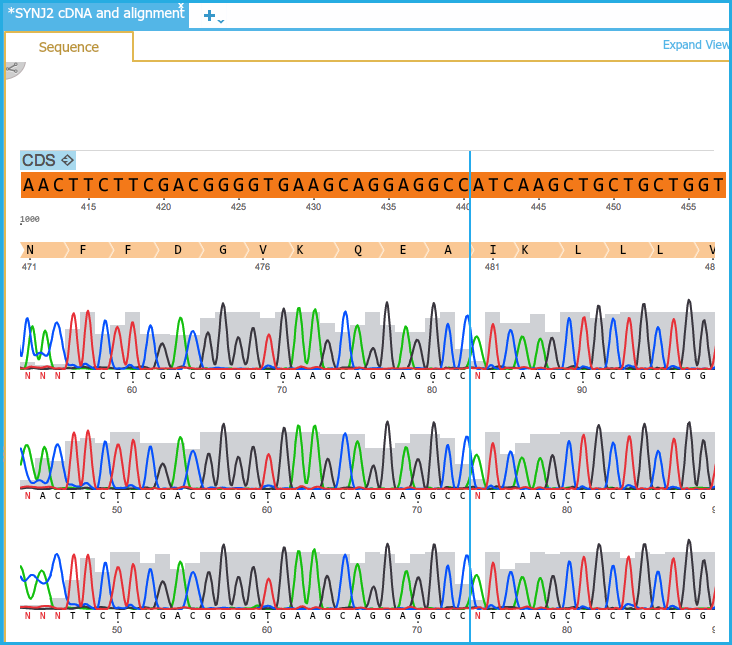 Figure 1.24.4.1: DNA View Alignments.
Figure 1.24.4.1: DNA View Alignments.</div>
Note: If the sequence for alignment does not contain chromatogram data then it will appear as a plain text sequence below the region in the project which it aligns to. add figure 12 (Figure 1.24.4.2).
 Figure 1.24.4.2: Plain Text DNA View.
Figure 1.24.4.2: Plain Text DNA View.</div>
Circular View and Linear View
- Alignments are displayed as an additional layer inside the plasmid in Circular view (Figure 1.24.4.3) and below the sequence in the Linear view (Figure 1.24.4.4). However trimmed portions of sequence are not shown.
Green regions indicate correct alignments and red regions indicate misalignments.
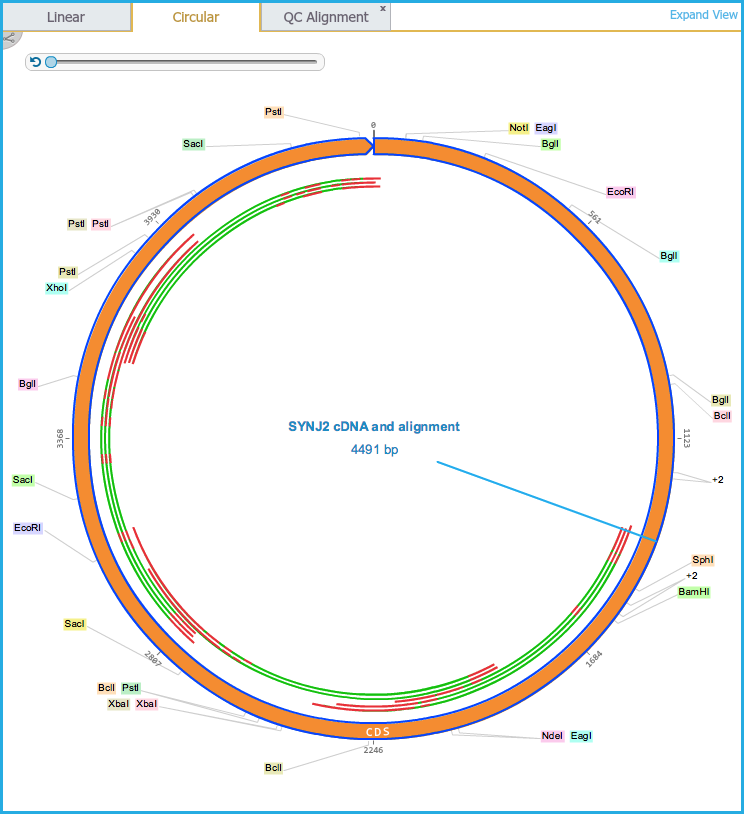 Figure 1.24.4.3: Circular View.
Figure 1.24.4.3: Circular View.</div>
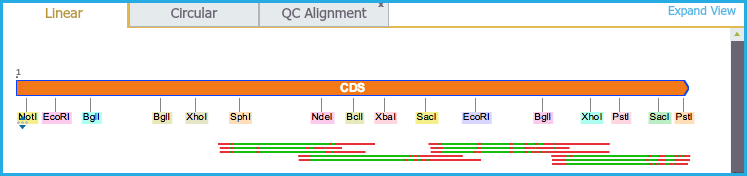 Figure 1.24.4.4: Linear View.
Figure 1.24.4.4: Linear View.</div>
Split Project View
If an alignment is selected in the Circular/Linear view the corresponding portion of sequence is selected in the Sequence view (Figures 1.24.4.5 and 1.24.4.6).
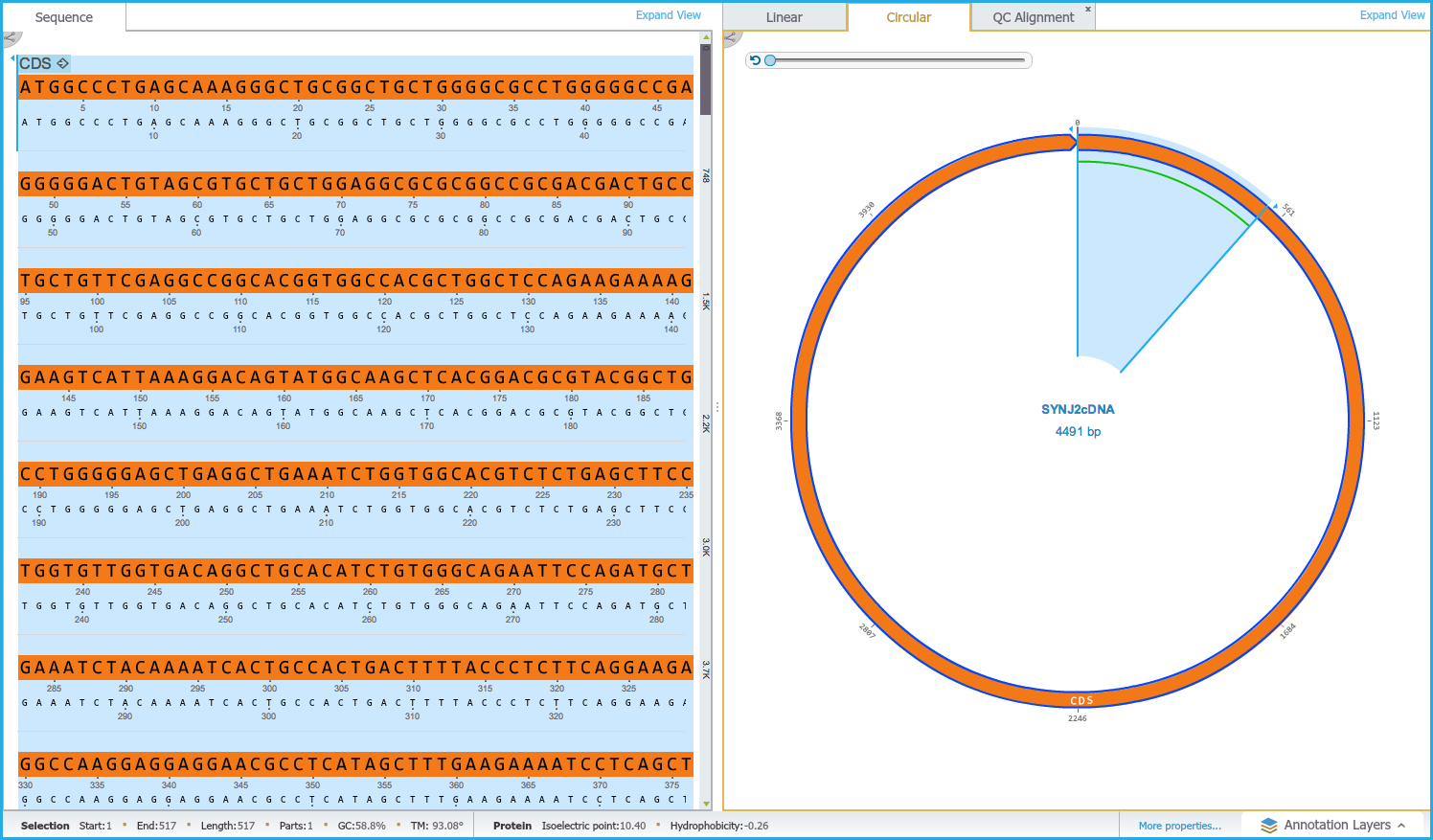 Figure 1.24.4.5: Dual View - Text Alignment.
Figure 1.24.4.5: Dual View - Text Alignment.</div>
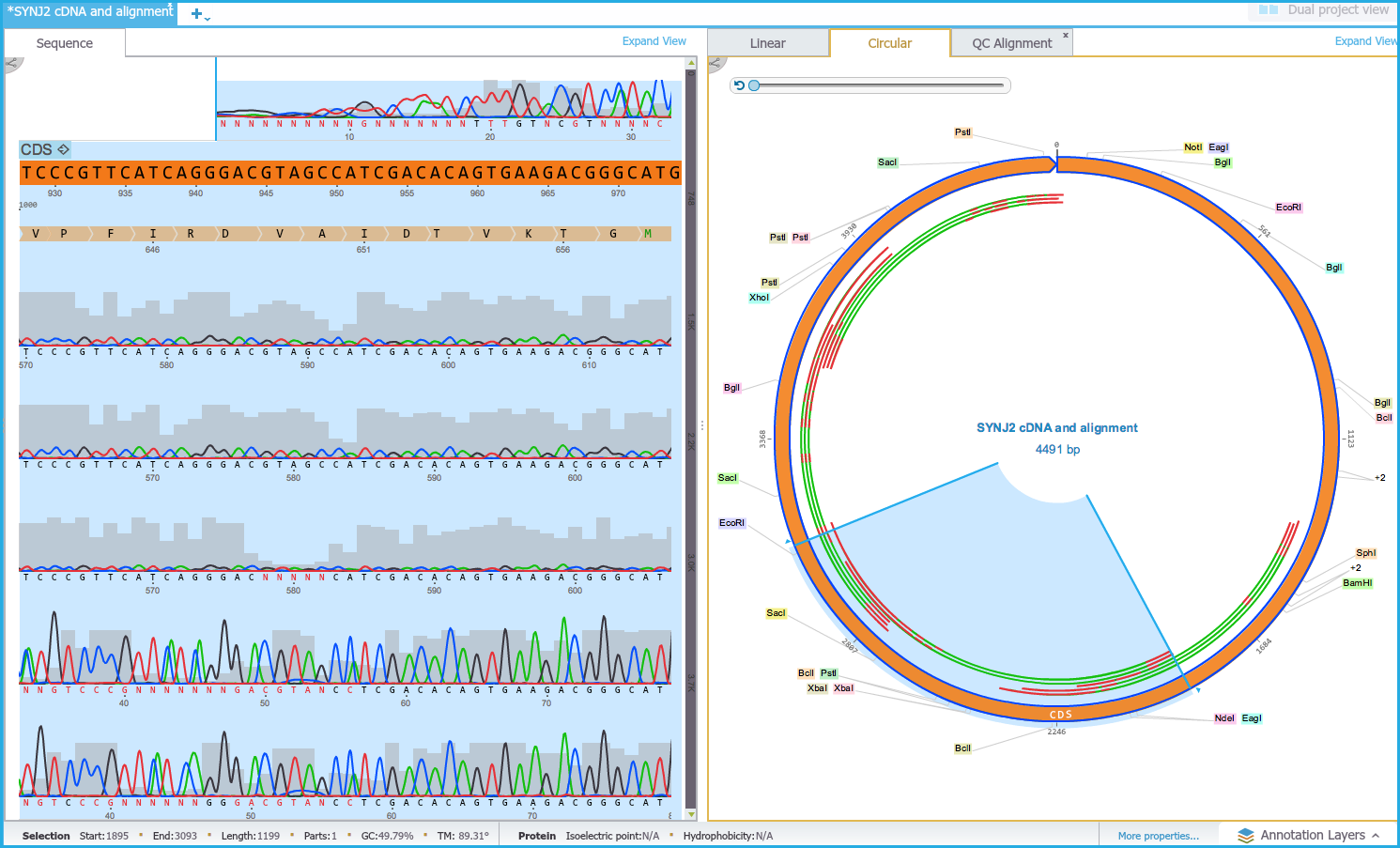 Figure 1.24.4.6: Dual View - Highlighting alignment.
Figure 1.24.4.6: Dual View - Highlighting alignment.</div>
Hiding the Alignment layer
You can hide the viewing of all QC Alignments by unchecking the "QC Alignment" box from the Annotation layers menu at the bottom right of the project (Figure 1.24.4.7).
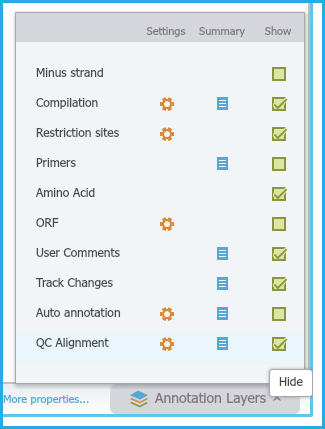 Figure 1.24.4.7: DNA View Alignments.
Figure 1.24.4.7: DNA View Alignments.</div>