1.19.3 Manual Primer Design Dialog
Please see the tutorial video below on "Manual Primer Design" for additional support:
The Manual Primer Design dialog (Figure 1.19.3.1) allows you to design, add descriptions, select a Primer Library to save to, and even pair your primers.
 Figure 1.19.3.1: The Manual Primer Design dialog.
Figure 1.19.3.1: The Manual Primer Design dialog.</div>
You can add a description and select the strand for your primer.
Choose "Select Library" to select a Primer Library to save the primer to.
To edit the primer, you can change the 5' location using the arrows, or select a different annealing sequence and click "Reset from selection" (Figure 1.19.3.2).
 Figure 1.19.3.2: The Manual Primer Design dialog: sequence ID, description, strand and 5' location.
Figure 1.19.3.2: The Manual Primer Design dialog: sequence ID, description, strand and 5' location.</div>
- In the "Overhang" section you can add Leader sequence, Restriction sites and Spacer sequences to your primers. (Figure 1.19.3.3).
You can open Genome Compilers codon wheel to help you choose which amino acid/s to add, and also select from a list of peptide coding sequences and restriction sites to add.
 Figure 1.19.3.3: Adding Leader and Spacer sequences and adding specific features.
Figure 1.19.3.3: Adding Leader and Spacer sequences and adding specific features.</div>
The Primer Actions section (Figure 1.19.3.1) allows you to create a copy of your primer, i.e. "Duplicate" it, and also to pair the primer with existing primers in your project. See section 1.19.5 for more details on how to pair primers.
The verification bar located at the bottom of the Primer dialog (Figure 1.19.3.5) displays the melting temperature (calculated according to [SantaLucia 1998] algorithm (http://www.pnas.org/content/95/4/1460.full)), the GC content and the length of your primers. Clicking "IDT Oligo Analyzer" will take you to the IDT website to check for secondary structure and hairpins formation in your primers.
The small red "x's" indicate that there is a basic primer design rule which has been violated. Hover on the "x" to see the design rule violation.
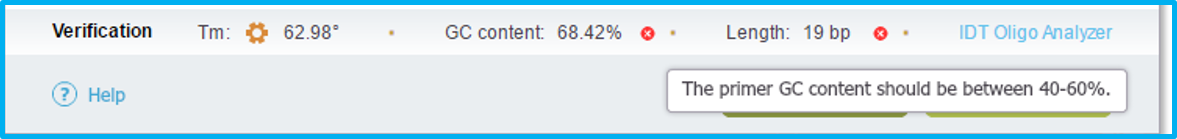 Figure 1.19.3.5: The Manual Primer Design dialog: verification.
Figure 1.19.3.5: The Manual Primer Design dialog: verification.</div>
- Clicking "Create" at the bottom of the primer dialog will save your primers to your chosen primer library and you can view the primers in the Primer Summary Table. See section 1.19.7 for more details on the Primer Summary Table.