1.23.4 Viewing ORFs
- ORFs appear as an additional annotation layer below the DNA sequence.
- ORFs in the same frame appear on the same row.
- Overlapping ORFs in different frames appear on different rows.
- Non-overlapping ORFs in the same frame or different frames appear in the same row.
An ORF sequence highlighted in the sequence view will be simultaneously selected in the adjacent circular/linear view and vice versa (Figure 1.23.4.1).
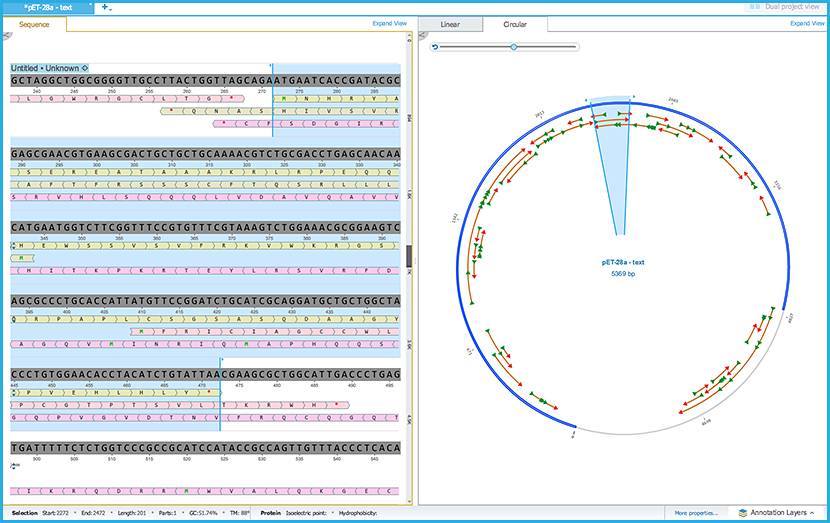 Figure 1.23.4.1: ORF Simultenous Viewing.
Figure 1.23.4.1: ORF Simultenous Viewing.</div>
Sequence View
- As in the amino acid layer, the directionality of each ORF is indicated with arrows along the ORF and stop codons are indicated with stars (*).
- Clicking anywhere in the ORF sequence will automatically select the nearest upstream start codon with its corresponding ORF sequence.
Clicking on a Methionine residue within an ORF sequence will automatically select specific sequence range within that ORF (Figure 1.23.4.2).
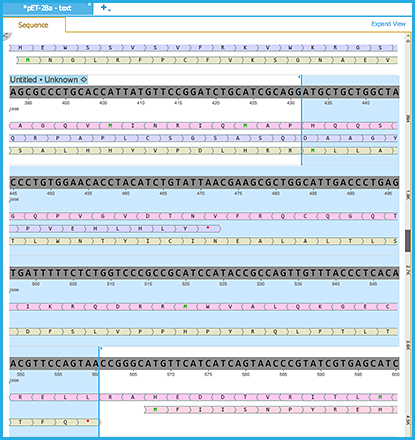 Figure 1.23.4.2: ORF Sequence View.
Figure 1.23.4.2: ORF Sequence View.</div>
Linear and Plasmid View
- ORFs are displayed in the same arrangement as the sequence view, however the directionality of each ORF is indicated with arrow heads.
- Each Methionine start codon is indicated with a green arrow head, whilst each stop codon is indicated with a red arrow head (according to the directionality of the part).
Clicking anywhere in the ORF sequence will automatically select the largest ORF sequence in that region (Figures 1.23.4.3 and 1.23.4.4).
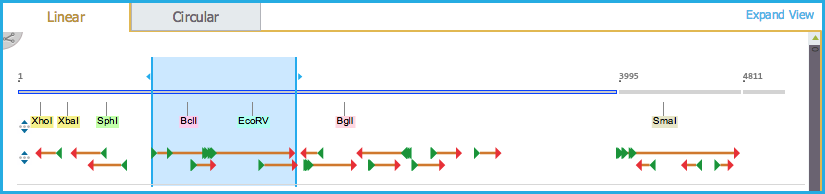 Figure 1.23.4.3: ORF Linear View.
Figure 1.23.4.3: ORF Linear View.</div>
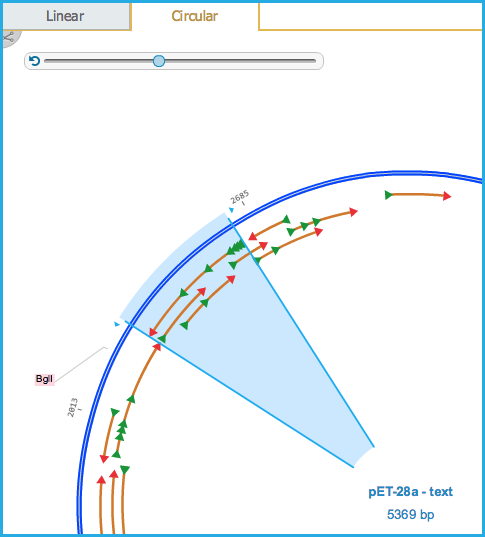 Figure 1.23.4.4: ORF Plasmid View.
Figure 1.23.4.4: ORF Plasmid View.</div>
Setting ORF as CDS
- This can be done from any view and allows you to set an ORF annotation as a CDS feature within the main layer of the sequence.
Select the ORF and then "Set as CDS" from the right click drop down menu (Figure 1.23.4.5).
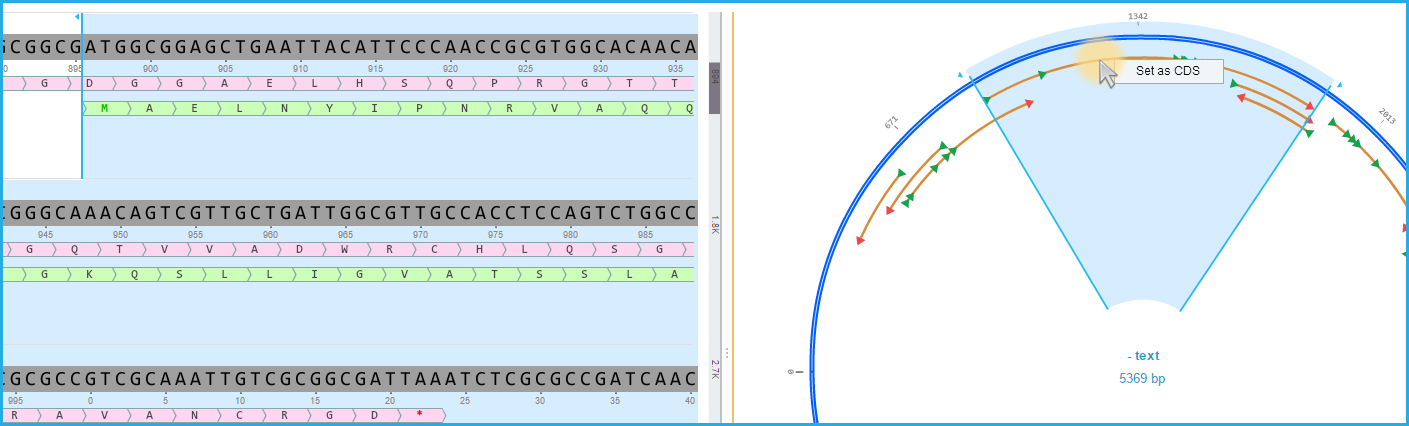 Figure 1.23.4.5: ORF set as CDS.
Figure 1.23.4.5: ORF set as CDS.</div>
The ORF will then be moved to the main layer and annotated as a CDS. Any pre-existing feature annotations will remain but will be split (Figure 1.23.4.6).
 Figure 1.23.4.6: CDS.
Figure 1.23.4.6: CDS.</div>